Objectives
Upon completion of this module topic, you will:
- be able to identify compatible restriction sites for cloning.
- be able to conduct a restriction digestion to prepare DNA fragments and vectors for the construction of recombinant DNA.
- be able to recombine a digested DNA fragment and plasmid vector in a ligation reaction to make recombinant plasmid DNA carrying a gene of interest.
- be able to perform a bacterial transformation reaction to transform competent bacterial cells with recombinant plasmid DNA.
- be able to make selective media plates for plating and growing transformed bacterial cells (antibiotic selective or X-gal).
- be able to plate transformed bacterial cells on selective plates and identifyresistant colonies.
- be able to screen resistant colonies for the gene of interest by amplifying individual bacterial colony DNA through Restriction Enzyme Digestion.
- be able to make an agarose gel, load the gel with your PCR reaction samples, and conduct agarose gel electrophoresis of samples.
- be able to isolate bacterial colonies carrying the plasmid by analyzing the agarose gel and determine samples containing the gene of interest.
- be able to make sterilized selective liquid media for growing up one of the bacterial colonies containing the gene of interest to produce large quantities of the desired plasmid DNA.
Part a
Constructing Recombinant DNA
This is Part A, Constructing Recombinant DNA, under the module topic, Molecular Cloning. This topic part has two sections: Content Tutorial & Animations.
DNA Cloning and Applications
Selection from the Campbell and Reece, AP Edition Biology, Seventh Edition, 2005 “Most methods for cloning pieces of DNA in the laboratory share certain general features. One common approach uses bacteria (most often, Escherichia coli) and their plasmids. Bacterial plasmids are relatively small, circular DNA molecules that replicate separate from a bacterial chromosome. For cloning genes or other pieces of DNA in the laboratory, a plasmid is first isolated from a bacterial cell, and then the foreign DNA is inserted into it. The resulting plasmid is now a recombinant DNA molecule, combining DNA from two sources. The plasmid is returned to a bacterial cell, producing a recombinant bacterium, which reproduces to form a clone of identical cells. Because the dividing bacteria replicate the recombinant plasmid and pass it on to their descendants, the foreign gene is “cloned” at the same time. Therefore the clone of cells contains multiple copies of the gene. Cloned genes are useful for two basic purposes: to make many copies of a particular gene and to produce a protein product. Researchers can isolate copies of a cloned gene from bacteria for use in basic research or to endow an organism with a new metabolic capability, such as pest resistance. For example, a resistance gene present in one crop species might be cloned and transferred into plants of another species. Alternatively, a protein with medical uses, such as human growth hormone, can be harvested in large quantities from bacterial cultures carrying the cloned gene for the protein. Most protein-coding genes exist in only one copy per genome—something on the order of one part per million of DNA—so the ability to clone such rare DNA fragments is extremely valuable.” (Campbell & Reece, AP Edition Biology Seventh Edition, pp. 385-386)
Technique Overview: Cloning a Human Gene in a Bacterial Plasmid
“Cloning is used to prepare many copies of a gene of interest for use in sequencing the gene, in producing its encoded protein, in gene therapy, or in basic research.” (Campbell & Reece, AP Edition Biology Seventh Edition, p.387) Real-Life Application of Molecular Cloning: Producing Human Insulin The following text and figure provides a preview into the steps involved in molecular cloning by providing a real-life application of molecular cloning techniques for the production of the human insulin protein. Read through the following text and examine the figure provided to see an overview of how to make recombinant DNA.
The Tools of Genetics: Recombinant DNA and Cloning – National Institute of General Medical Sciences, NIH
- Recombinant DNA and Cloning (publications.nigms.nih.gov, HTML Page)
“In the early 1970s, scientists discovered that they could change an organism’s genetic traits by putting genetic material from another organism into its cells. This discovery, which caused quite a stir, paved the way for many extraordinary accomplishments in medical research that have occurred over the past 35 years.

Scientists in Scotland were the first to clone an animal, this sheep named Dolly. She later gave birth to Bonnie, the lamb next to her. ROSLIN INSTITUTE, EDINBURGH
How do scientists move genes from one organism to another? The cutting and pasting gets done with chemical scissors: enzymes, to be specific. Take insulin, for example. Let’s say a scientist wants to make large quantities of this protein to treat diabetes. She decides to transfer the human gene for insulin into a bacterium, Escherichia coli, or E. coli, which is commonly used for genetic research. That’s because E. coli reproduces really fast, so after one bacterium gets the human insulin gene, it doesn’t take much time to grow millions of bacteria that contain the gene. The first step is to cut the insulin gene out of a copied, or “cloned,” version of the human DNA using a special bacterial enzyme from bacteria called a restriction endonuclease. (The normal role of these enzymes in bacteria is to chew up the DNA of viruses and other invaders.) Each restriction enzyme recognizes and cuts at a different nucleotide sequence, so it’s possible to be very precise about DNA cutting by selecting one of several hundred of these enzymes that cuts at the desired sequence. Most restriction endonucleases make slightly staggered incisions, resulting in “sticky ends,” out of which one strand protrudes. The next step in this example is to splice, or paste, the human insulin gene into a circle of bacterial DNA called a plasmid. Attaching the cut ends together is done with a different enzyme (obtained from a virus), called DNA ligase. The sticky ends join back together kind of like jigsaw puzzle pieces. The result: a cut-and-pasted mixture of human and bacterial DNA. The last step is putting the new recombinant DNA see definition (publications.nigms.nih.gov, HTML Page) back into E. coli and letting the bacteria reproduce in a petri dish. Now, the scientist has a great tool: a version of E. coli that produces lots of human insulin that can be used for treating people with diabetes. So, what is cloning? Strictly speaking, it’s making many copies of a gene—in the example below, E. coli is doing the cloning. However, the term cloning is more generally used to refer to the entire process of isolating and manipulating a gene. Dolly the cloned sheep contained the identical genetic material of another sheep. Thus, researchers refer to Dolly as a clone (publications.nigms.nih.gov, HTML Page). clone #2(publications.nigms.nih.gov, HTML Page)
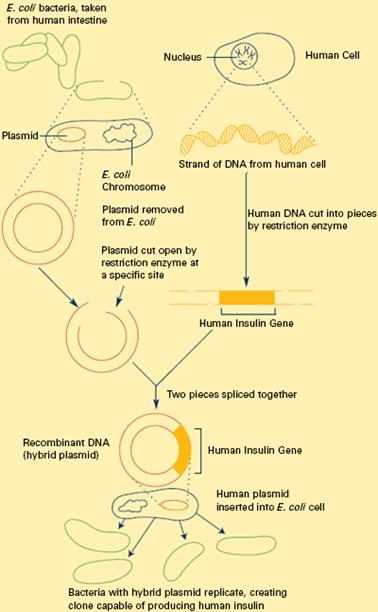
Constructing Recombinant DNA: Above Figure To splice a human gene (in this case, the one for insulin) into a plasmid, scientists take the plasmid out of an E. coli bacterium, cut the plasmid with a restriction enzyme, and splice in insulin-making human DNA. The resulting hybrid plasmid can be inserted into another E. coli bacterium, where it multiplies along with the bacterium. There, it can produce large quantities of insulin.
Cloning Vectors
From The College Board AP Biology Lab Manual, Student Edition (2001) pp.64-65. “The bacterium Escherichia coli (E. coli) is an ideal organism for the molecular geneticist to manipulate and has been used extensively in recombinant DNA research. It is a common inhabitant of the human colon and can easily be grown in suspension culture in a nutrient medium such as Luria broth, or in a petri dish of Luria broth mixed with agar (LB agar) or nutrient agar.
The single circular chromosome of E. coli contains about five million DNA base pairs, only 1/600th the haploid amount of DNA in a human cell. In addition, the E. coli cell may contain small, circular DNA molecules (1,000 to 200,000 base pairs called plasmids, which also carry genetic information. The plasmids are extrachromosomal; they exist separately from the chromosome. Some plasmids replicate only when the bacterial chromosome replicates and usually exist only as single copies within the bacterial cell. Others replicate autonomously and often occur in as many as 10 to 200 copies within a single bacterial cell. Certain plasmids called R plasmids, carry genes for resistance to antibiotics such as ampicillin, kanamycin, or tetracycline.
In nature genes can be transferred between bacteria in three ways: conjugation, transduction, or transformation. Conjugation is a mating process during which genetic material is transferred from one bacterium to another of a different mating type. Transduction requires the presence of a virus to act as a vector (carrier) to transfer small pieces of DNA from one bacterium to another. Bacterial transformation involves transfer of genetic information into a cell by direct uptake of the DNA. During gene transfer, the uptake and expression of foreign DNA by a recipient bacterium can result in conferring a particular trait to a recipient lacking that trait. Transformation can occur naturally but the incidence is extremely low and is limited to relatively few bacterial strains. These bacterial can take up DNA only during the period at the end of logarithmic growth. At this time the cells are said to be competent. Competence can be induced in E.coli with carefully controlled chemical growth conditions. Once competent, the cells are ready to accept DNA that is introduced from another source.
Plasmids can transfer genes (such as those for antibiotic resistance) that occur naturally within them, or plasmids can act as carriers (vectors) for introducing foreign DNA from other bacteria, plasmids, or even eukaryotes (animals, plants, fungi, protists) into bacterial cells. Restriction endonucleases (enzymes) can be used to cut and insert pieces of foreign DNA into the plasmid vectors. If these plasmid vectors also carry genes for antibiotic resistance, transformed cells containing plasmids that carry the foreign DNA of interest in addition to the antibiotic resistance gene can be easily selected from other cells that do not carry the gene for antibiotic resistance.” (College Board AP Biology Lab Manual, Student Edition (2001) pp.64-65)
Plasmid Maps
The following link will allow you to view three plasmid maps and their respective nucleotide sequences. The lowercase “p” stands for plasmid and the three maps are for “R” plasmids (resistance): pAMP (plasmid with an ampicillin resistance gene), pKAN (plasmid with kanamycin resistance gene), pBLU (plasmid with ampicillin resistance gene and Lac Z gene for lactose breakdown- encodes β-galactosidase). Each plasmid map also indicates the size of the plasmid in DNA base pairs. The restriction sites for specific restriction enzymes are annotated on each plasmid map at the specific location in which the particular restriction enzyme indicated (for example: EcoRI, HindIII, BamHI, etc.) will cut the plasmid. When you click on the plasmid map you will be able to view the entire sequence of each plasmid. Task: Click on the following link to view the following plasmid maps and sequences. Cold Spring Harbor Laboratory: Nucleotide Sequences of Plasmids
- Plasmid Maps (www.dnalc.org, Multimedia Page)
Animations
1. Plasmid Animation: The Ti Plasmid Animation from Raven, Johnson, Losos, & Singer, Biology, 7th Edition, 2005 Click on the following link to view the animation list. Click on the animation titled “Ti Plasmid” to learn about the Ti plasmid, a useful vector for the genetic manipulation of plants.
- The Ti Plasmid (highered.mcgraw-hill.com, Multimedia Page)
2. Plasmid Applications: Constructing Vaccines Animation from Raven, Johnson, Losos, & Singer, Biology, 7th Edition, 2005 Click on the following link to again view the animation list. Now click on the animation titled “Constructing Vaccines” to learn about the important role and medical applications that plasmids have in the construction of vaccines.
- Constructing Vaccines (highered.mcgraw-hill.com, Multimedia Page)
3. Plasmid Cloning: Sumanas Inc. Animation from Lodish, et al., Molecular Cell Biology, Fifth Edition,
- W. H. Freeman & Co. (2004) (www.whfreeman.com, Multimedia Page)
Click on the following link to learn about plasmid cloning. You will be provided with the options to view the animation in a narrated format or in a self-guided format with text explanations.
- Plasmid Cloning (www.sumansinc.com, Multimedia Page)
4. Plasmid Cloning Video: Biosolutions Click on the following link to view an animation that depicts the steps involved in cloning a plasmid of interest. This animation also applies to the sections to follow in this module subset because the cloning process involves all of these steps including, restriction enzyme digestion, DNA ligation, and bacterial transformation.
- Plasmid Cloning Video (bioisolutions.blogspot.com, Multimedia Page)
5. Steps in Cloning a Gene Animation from Raven, Johnson, Losos, & Singer, Biology, 7th Edition, 2005 View the following narrated animation to see a simulated overview about the various laboratory steps involved in cloning a gene. The steps will be broken down and examined in greater detail in the following module subset sections. The animation serves to provide an integrated overview of the sequence of events. When you click on the link below you will be taken to a list of molecular biology Animations. View the narrated animation titled “Steps in Cloning a Gene”.
- Steps in Cloning a Gene (highered.mcgraw-hill.com, Multimedia Page)
6. Real-Life Medical Applications – Explore how recombinant DNA technology is used to make synthetic insulin. Click on the following link provided below to view the “Manipulation” section from the menu of the Cold Spring Harbor Laboratories DNA Interactive Website. Click on the “Production” module. It is not necessary to move from one section to another in sequence. The “putting it together” section allows you to view animations about how synthetic insulin was made using recombinant DNA through both bacterial and yeast methods. After viewing the animations you may view “The Genentech Tour” video below the animations to learn about how the first biotechnology company, Genentech, was started in 1976. “Today, Genentech makes and holds the patents for a number of products used in the treatment of medical conditions”
- Production (www.dnai.org, Multimedia Page) Click on Production, then click on Putting it Together
- The Genentech Tour (www.dnai.org, Multimedia Page) Click on Production, click on Putting it Together, then click on The Genentech Tour
Part b
Restriction Enzyme Digestion
This is Part B, Restriction Enzyme Digestion, under the module topic, Molecular Cloning. This topic part has two sections: Content Tutorial and Animations.
Content Tutorial
Restriction Enzymes and Cloning
This section on cloning includes information that is a necessary basic for research laboratories, but may be required from time to time in clinical labs when something really interesting is found in the molecular lab. Cloning of a gene sequence (or part) is also required to have stable templates for the production of probes, or to save that interesting PCR product. Long term storage of PCR products without cloning them, leaves them susceptible to degradation by bacterial exonucleases (enzymes that digest DNA from the ends). Restriction enzymes are the backbone reagents of cloning, but are used in clinical applications associated with fingerprinting – genetic identity, epidemiology, and in preparation for blotting for other applications.
Cloning Step 1: Restriction digestion.
This is a step that essentially cuts DNA into little bits. Recall from module subset II-a, that there was also material on restriction enzyme digestions that has provided some background for this section and also included the application of restriction digestions to identification (DNA fingerprinting). The molecular scissors are called restriction endonucleases. Recall that restriction endonucleases are enzymes that cleave DNA. They are derived from bacteria where they function in cleaving foreign DNA and thus protecting the integrity of the host bacteria (a bacterial immune system). These enzymes recognize specific base sequences in double stranded DNA. The base sequence serves as a code for the restriction enzyme saying “cut here”. For example, the enzyme labelled EcoRI recognizes the GAATTC sequence of bases and cuts between the G and A.
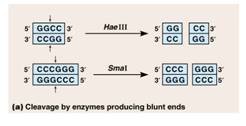
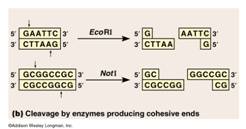
AWLInc
The sites on the DNA molecule recognized by the enzymes are called restriction sites. Restrictions sites are palindromic, reading the same 5’→3’ on either strand of the DNA. The various enzymes are named for the bacteria from which they have been derived. Eco is derived from Escherichia coli and Hin from Haemophilus influenzae, for example. You will note that the first panel above the cut leaves no overhang and is called a blunt cut leaving blunt ends. Any blunt end can be rejoined to any other blunt end – something to remember when cloning PCR products. The other panel shows a cut that leaves overhanging ends called either 5’ or 3’ overhangs depending on which strand is overhanging.
Enzymes may be chosen that will cleave on either side of a desired DNA sequence. In these cases, cut sites have been predetermined by finding the sequence of nucleotides that encode a specific protein or characteristic. In other cases, various endonucleases will be tried.
Once the DNA has undergone restriction digestion, it may be used to recombine with any other piece of DNA that has the complementary ends, regardless of the source of that DNA. This allows insertion of a segment of DNA into a plasmid that has been cleaved with the same enzyme(s)
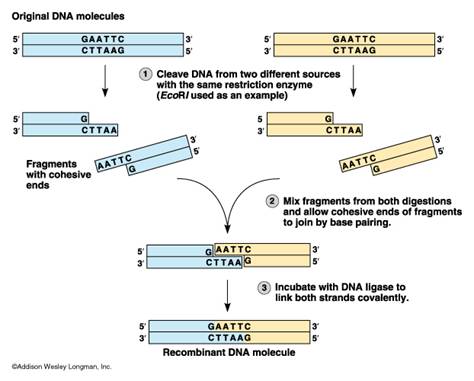
University of Calgary Biotechnology Training Centre
Animations
1. Restriction Endonucleases Animation from Raven, Johnson, Losos, & Singer, Biology, 7th Edition, 2005 Click on the following link to view the animation list. Click on the animation titled “Restriction Endonucleases” to learn about restriction enzymes and their applications to molecular cloning and making recombinant DNA.
- Restriction Endonucleases (highered.mcgraw-hill, Multimedia Page)
2. Plasmid Applications: Constructing Vaccines Animations from Raven, Johnson, Losos, & Singer, Biology, 7th Edition, 2005 Click on the following link to again view the animation list. Now click on the animation titled “Constructing Vaccines”. This animation was provided in the Constructing Recombinant DNA section and it is provided again for reinforcement. When you view the animation this time, focus on the role that restriction enzymes play in constructing vaccines.
- Constructing Vaccines (highered.mcgraw-hill, Multimedia Page)
Part c
DNA Ligation
This is Part C, DNA Ligation, under the module topic, Molecular Cloning. This topic part has two sections: Content Tutorial and Animations
Content Tutorial
Cloning Step 2: The Vector and Ligation Plasmids are circular, extrachromosomal bits of double stranded DNA found in some bacteria. Plasmids use the bacterial cells energy and metabolic pathways to replicate themselves. For a plasmid to be useful as a vector, it should be reasonably small for easy transfer into bacterial cells and be capable of replicating itself in large numbers.
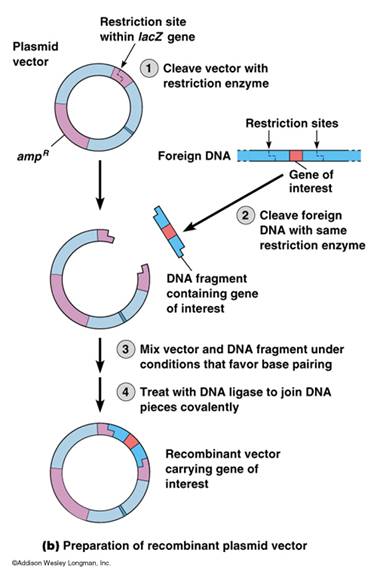
A plasmid must be selected with a restriction site identical to the restriction sites in the foreign DNA. If the plasmid and foreign DNA are treated with the same restriction endonuclease, the “sticky ends,” or ends with 5’ or 3’ overhangs, of the foreign DNA will stick or reanneal to the sticky ends of the plasmids. Thus the DNA fragment is inserted into the plasmid vector. A second enzyme, DNA ligase, repairs the breaks in the strands. DNA Ligase is an enzyme required during DNA replication to fix the breaks in DNA and as part of our DNA repair mechanism.
As previously mentioned in module subset II-b, PCR amplicons can be cloned directly with out digesting the ends, but often generating “sticky ends” is more successful. Such ends must be planned to be included in the primers (see PCR section in Nucleic Acid Sequencing).
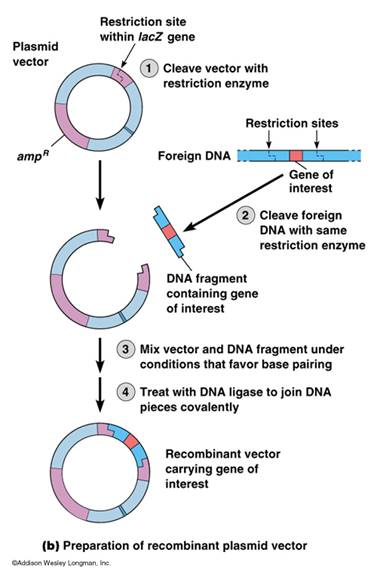
Construction of a Recombinant Cloning Vector – University of Calgary Biotechnology Training Center
Animations
1. Restriction Endonucleases Animation from Raven, Johnson, Losos, & Singer, Biology, 7th Edition, 2005 Click on the following link to view the animation list. Click on the animation titled “Restriction Endonucleases”. This animation was provided in the previous section and it is provided again for reinforcement. When you view the animation this time, focus on the role that DNA ligase plays in the ligation reaction.
- Restriction Endonucleases (highered.mcgraw-hill.com, Multimedia Page)
Part d
Bacterial Transformation
This is Part D, Bacterial Transformation, under the module topic, Molecular Cloning. This topic part has three sections: Content Tutorial, Animations, and Activities.
Content Tutorial
Cloning Step 3: Transformation
The plasmid containing the foreign DNA is called a plasmid chimera. The plasmid chimeras are then mixed with an organism such as Escherichia coli in the presence of calcium phosphate or calcium chloride. This alters the membrane permeability of the E. coli and facilitates uptake of the plasmid. The plasmid then propagates in the host bacterial cell and produces many copies of the foreign DNA.
Not all the ligated vectors will contain an insert. In addition, not all the bacteria exposed to the chimeric plasmid will take up the plasmid. Techniques have been developed to select the desired colonies.
A plasmid commonly used carries the genes coding for antibiotic resistance, such as ampicillin or tetracycline resistance. These genes are called selection markers. Another common selection uses one of the lacZ genes from bacteria that encodes the β-galactosidase enzyme. β-gal activity is detected by a substrate X-gal (a lactose analog) that turns blue when hydrolyzed. By using the right restriction endonuclease, the plasmid can be cleaved through the site coding for β-galactosidase. When the foreign DNA is inserted, the gene for the β-galactosidase is no longer functional.
In the diagram below, the chimeric plasmid carries resistance to ampicillin but the β-galactosidase gene has been disrupted. The bacteria are grown on a solid medium with ampicllin and X-gal. Cells resistant to ampicillin grow into colonies and white colonies (lac-) are those carrying the chimeric plasmid with the foreign DNA. Those specific colonies are grown, harvested, and stored. Plasmids isolated from subcultures can be used for sequencing the insert, storage, RFLP analysis or the preparation of a probe.
University of Calgary Biotechnology Training Center
Animations
1. 3D Animation: Genetic Engineering The following 3D Animation depicts the steps involved in molecular cloning providing a visual model that integrates Molecular Cloning (II-d) Sections Video: 3-D Animation: Genetic Engineering
- Cloning Vectors
- Restriction Digestion
- DNA Ligation with this current section
- Bacterial Transformation
The following animation reviews the role of restriction enzymes in gene cloning and genetic engineering (Module Subset II.a.). The animation portrays the sequence of events involved in making recombinant DNA by displaying a bacterial plasmid undergoing a restriction digestion with restriction enzymes in addition to the insertion and ligation of a new DNA segment into the original plasmid. The bacterial cell will then undergo transformation to produce clones of the new recombinant DNA and can produce the proteins encoded by the newly inserted gene.
- Genetic Engineering (www.hhmi.org, Multimedia Page)
2. Biology Animation Library: DNA Transformation “Stanley Cohen and Herbert Boyer’s historic experiment used techniques to cut and paste DNA to create the first custom-made organism containing recombined or “recombinant” DNA. Cohen and Boyer inserted the recombinant DNA molecule they created into E. coli bacteria by means of a plasmid, thereby inducing the uptake and expression of a foreign DNA sequence known as “transformation.” Dolan DNA Learning Center, DNA InteractiveTask:
- View Animation 1: (www.dnalc.org, Multimedia Page)
DNA Transformation Continued: “DNA transformation is a naturally occurring but rare event in which DNA can be transferred into bacteria. In 1970, Morton Mandel and Akiko Higa discovered a way to make E. coli more “competent” for transforming foreign DNA. Their calcium chloride method is widely used today to obtain high-efficiency transforming cells.”
Dolan DNA Learning Center, DNA Interactive
Task:
- View Animation 2: (www.dnalc.org, Multimedia Page)
Activities
1. Lab Bench Activity: Molecular Biology Lab 6-I Bacterial Transformation
- Bacterial Transformation (www.phschool.com, Multimedia Page)
“In this laboratory you will use some basic tools of molecular biology to gain an understanding of some of the principles and techniques of genetic engineering. In the first part of the lab, you will use antibiotic-resistance plasmids to transform Escherichia coli. In the second part, you will use gel electrophoresis to separate fragments of DNA for further analysis.”
- Lab Introduction (www.phschool.com, Multimedia Page)
*Note: If following the subsets in order you will have already completed the Second Part of this activity in Module Subset II-a, Section 2. Task: Complete Lab Bench Activity 6-I. Bacterial Transformation, Analysis of Results, and Lab Quiz 6-I. Record your analysis of Results I and Lab Quiz I.
- Bacterial Transformation (www.phschool.com, Multimedia Page)
2. AP Biology Virtual Lab: Bacterial Transformation University of California College Prep Open Access Materials (UCCP Open Access) Task: The following website provided below contains a virtual molecular biology lab for conducting a bacterial transformation experiment. When you access the webpage select Lab 8, Molecular Biology. When you are viewing the lab homepage you can begin by reviewing the “getting started…” materials that include an overview of the lab, video, lab manual, equipment, and procedure demonstration. Once you have finished going through the pre-lab materials, click on the arrow to enter the “sim lab”, where you will be directed to your virtual lab bench where you will be conducting the experiment. On the right side of your lab bench screen you are provided with a virtual stock room where you will find your laboratory materials and trash for waste disposal. On the left hand side of your virtual lab bench screen you are provided with access to the original “getting started” materials that you will be able to reference at any time during your experiment.
- Virtual Lab:Bacterial Transformation (www.ucopenaccess.org, Multimedia Page)
3. Case Study: Applied Genetic Modification Annenberg/CPB Media Click on the following link to access the Case Study, “Applied Genetic Modification”. You will be presented with an introduction to the case study and some recommended materials to view before beginning including a Genetically Modified Organisms Video and Online Text reading. When you have finished going through the introduction materials, click on the green “Go” arrow to “Launch Case Study”. (Both “Flash” and “HTML” versions are available on the website for viewing). The Case Study is organized into the following sections:
- Case Study (www.learner.org, HTML Page)
Overview
- Select a Plant Delivery System
- Select a Viral Antigen
- Overview of Cloning: Major Steps Involved in Cloning a Gene
- Screening of a Library
- Features of a Plasmid Vector for Making Transgenic Plants: Typical Bacteriophage Vector Used for DNA Ligation
- Ligating a Gene Into a Plasmid
- Plasmid Mapping: Colonies of e.coli bacteria with the inserted plasmid
- Plasmid Mapping: Restriction Enzyme Digest
- Plasmid Mapping: Gel Electrophoresis
- Transformation Methods
- Expression Data on Transgenic Plants: Northern Blot- RNA Levels (review of Nucleic Acid Hybridization). (*Note: The Western Blot- Protein Levels applies to Module III Protein Techniques)
- Vaccine Development – Antibody Detection and Measurement (applies to Protein Techniques)
4. Arizona Biology Project: Recombinant DNA Technology Problem Set In this problem set, you will learn about some of the basic techniques of recombinant DNA, and how recombinant DNA technology is applied to human health. Task: Complete problems 2, 4, 5, 9, 10, & 11 from the set. The following problems will have multiple choice answers. Correct answers are reinforced with a brief explanation. Incorrect answers are linked to tutorials to help solve the problem.
- Recombinant DNA Technology Problem Set (www.biology.arizona.edu, Multimedia Page)
5. Extended Practice: Recombinant DNA and Cloning Practice Problems & Solutions MIT AP Biology Open Courseware
- Practice Problem 1: (ocw.mit.edu, HTML Page)
- Solution: (ocw.mit.edu, HTML Page)
- Practice Problem 2:(ocw.mit.edu, HTML Page)
- Solution:(ocw.mit.edu, HTML Page)
Part e
DNA and Gene Libraries
This is Part E, DNA and Gene Libraries, under the module topic, Molecular Cloning. This topic part has two sections: Content Tutorial andAnimations.
Content Tutorial
Storing Cloned Genes in DNA Libraries “A molecular cloning procedure that starts with a mixture of fragments from the entire genome of an organism, is called a “shotgun” approach; no single gene is targeted for cloning. Thousands of different recombinant plasmids are produced and a clone of each ends up at the end of the procedure. The complete set of plasmid clones, each carrying copies of a particular segment from the initial genome, is referred to as a genomic library. Scientists often obtain such libraries (or even particular cloned genes) from another researcher or a commercial source (sometimes referred to as “cloning by phone”!). Certain bacteriophages are also common cloning vectors for making genomic libraries. Fragments of foreign DNA can be spliced into a phage genome, as into a plasmid, by using a restriction enzyme and DNA ligase. An advantage of using phages as vectors is that a phage can carry a larger DNA insert than a bacterial plasmid. The recombinant phage DNA is packaged into capsids in vitro and introduced into a bacterial cell through the normal infection process. Inside the cell, the phage DNA replicates and produces new phage particles, each carrying the foreign DNA. A genomic library made using phage is stored as a collection of phage clones. Since restriction enzymes do not recognize gene boundaries, some genes in either type of genomic library will be cut and divided up among two or more clones. Researchers can make another kind of DNA library starting with mRNA extracted from cells. The enzyme reverse transcriptase (obtained from retroviruses) is used in vitro to make single–stranded DNA transcripts of the mRNA molecules. Following enzymatic degradation of the mRNA, a second DNA strand, complementary to the first, is synthesized by DNA polymerase. This double–stranded DNA, called complementary DNA (cDNA), is then modified by addition of restriction enzyme recognition sequences at each end. Finally, the cDNA is inserted into vector DNA in a manner similar to the insertion of genomic DNA fragments. The isolated mRNA was a mixture of all the mRNA molecules in the cells used, transcribed from many different genes. Therefore, the cDNAs that are cloned make up a library containing a collection of genes. However, a cDNA library represents only part of the genome—only the subset of genes that were transcribed into mRNA in the original cells. Genomic and cDNA libraries each have advantages, depending on what is being studied. If you want to clone a gene but are unsure in what cell type it is expressed or are unable to obtain that cell type, a genomic library is almost certain to contain the gene. Also, if you are interested in the regulatory sequences or introns associated with a gene, a genomic library is required because these sequences are absent from the fully processed mRNAs used in making a cDNA library. For this very reason, if you are only interested in the coding sequence of a gene, you can obtain a stripped–down version of the gene from a cDNA library. A cDNA library is also useful for studying genes responsible for the specialized functions of a particular cell type, such as brain or liver cells. Finally, changes in patterns of gene expression during development can be traced by making cDNA from cells of the same type at different times in the life of an organism.” (Campbell & Reece, AP Edition Biology Seventh Edition, pp. 387-388)
Question: What is the difference between a genomic DNA library and a cDNA library? Identify some examples of situations in which it would be more advantages to use one over the other for studying genes.
CONTENT TUTORIAL 2: Lodish et al., Molecular Cell Biology, W H Freeman and Company (2000)
The following link provides a tutorial in which you will learn about the molecular techniques used for constructing DNA libraries with various cloning vectors. There is a summary at the end of the tutorial highlighting the major concepts for the techniques involved. There are seven figures that go along with the tutorial that are helpful for depicting the techniques explained, you can access the figures by clicking on their links on the right hand side of the tutorial webpage. For example, one very useful figure is figure 7-12. This figure portrays the steps involved in constructing a genomic library of human DNA in a bacteriophage (virus that infects bacteria) λ vector.
- Constructing DNA libraries with Various Cloning Vectors (www.ncbi.nlm.nih.gov, Multimedia Page)
Selection from the Campbell and Reece, AP Edition Biology, Seventh Edition, 2005
Animations
1.“Storing DNA” Cold Spring Harbor Laboratory DNA Interactive
a. Click on the link provided below to view the “Manipulation” section from the menu of the Cold Spring Harbor Laboratory’s DNA Interactive Website. Next, click on the “Techniques” module and view the “Transferring & Storing” section. The “Transferring & Storing” technique section will allow you to review DNA Transformation or you can move along to the Storing DNA 2D Animation to learn about the different types of DNA libraries (Yeast artificial chromosomes, Bacterial artificial chromosomes, Cosmids, Bacteriophage lambda, and plasmids). These sections also provide a good review of some of the various types of cloning vectors used in molecular cloning.
- Techniques (www.dnai.org, Multimedia Page)
Click on Techniques, then click on Transferring & Storage
b. When you have finished viewing the Storing DNA animation you may proceed to view the optional video “Interviews” provided to learn about the natural genetic engineering abilities of agrobacterium tumefaciens.
2. “Creating a DNA Library” Cain, et al., Discover Biology, Third Edition W. W. Norton & Co., 2006 W. W. Norton & Co. and Sumanas, Inc. The following website provides a multimedia tutorial and animation that you can go through in either a step-through or narrated format to learn about how researchers construct DNA libraries.
- Creating a DNA Library (www.sumanasinc.com, Multimedia Page)
When you are finished viewing the tutorial and animation, take the quiz by clicking on the Intro link, then click on Quiz